|
Introduction:
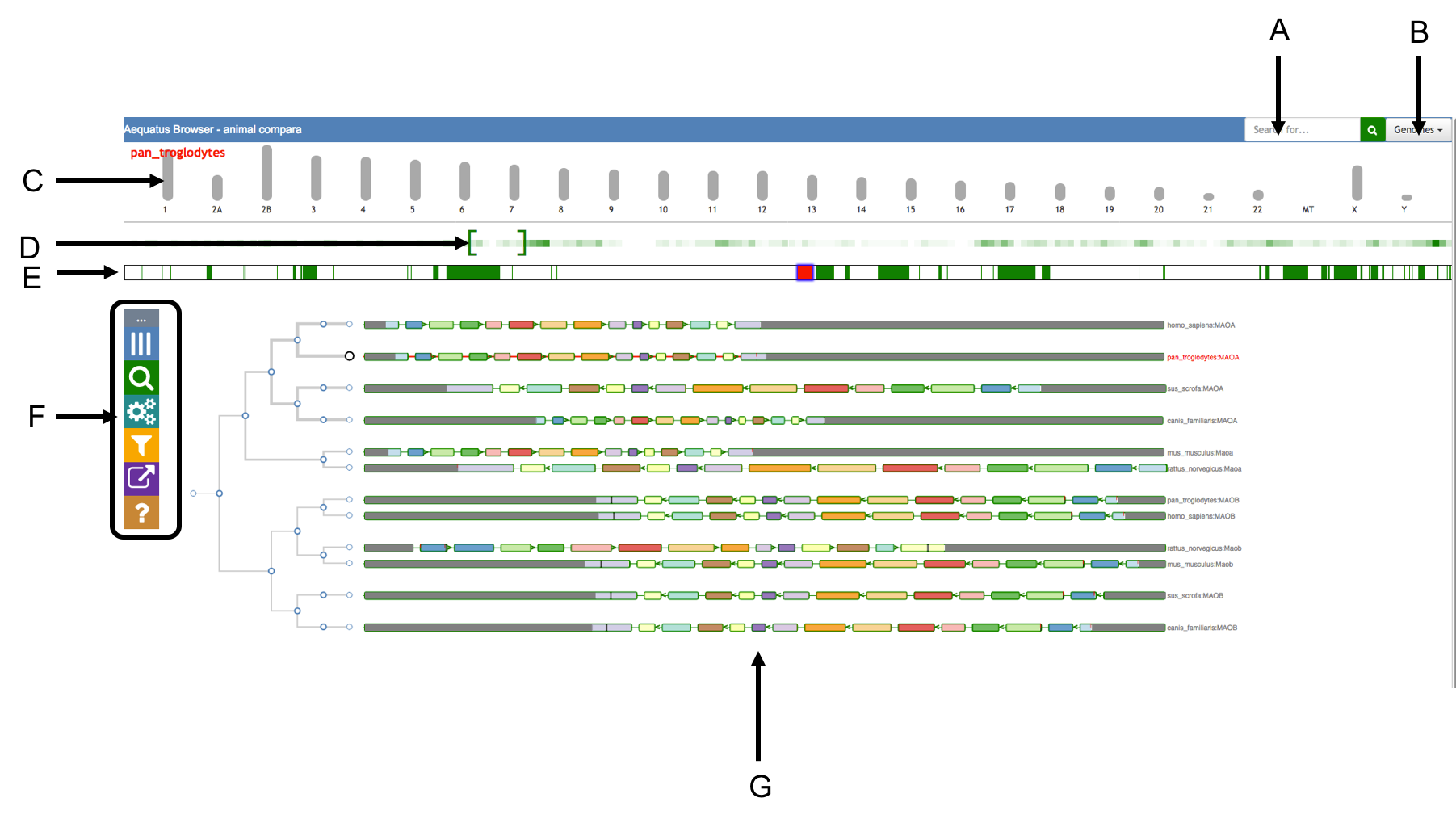
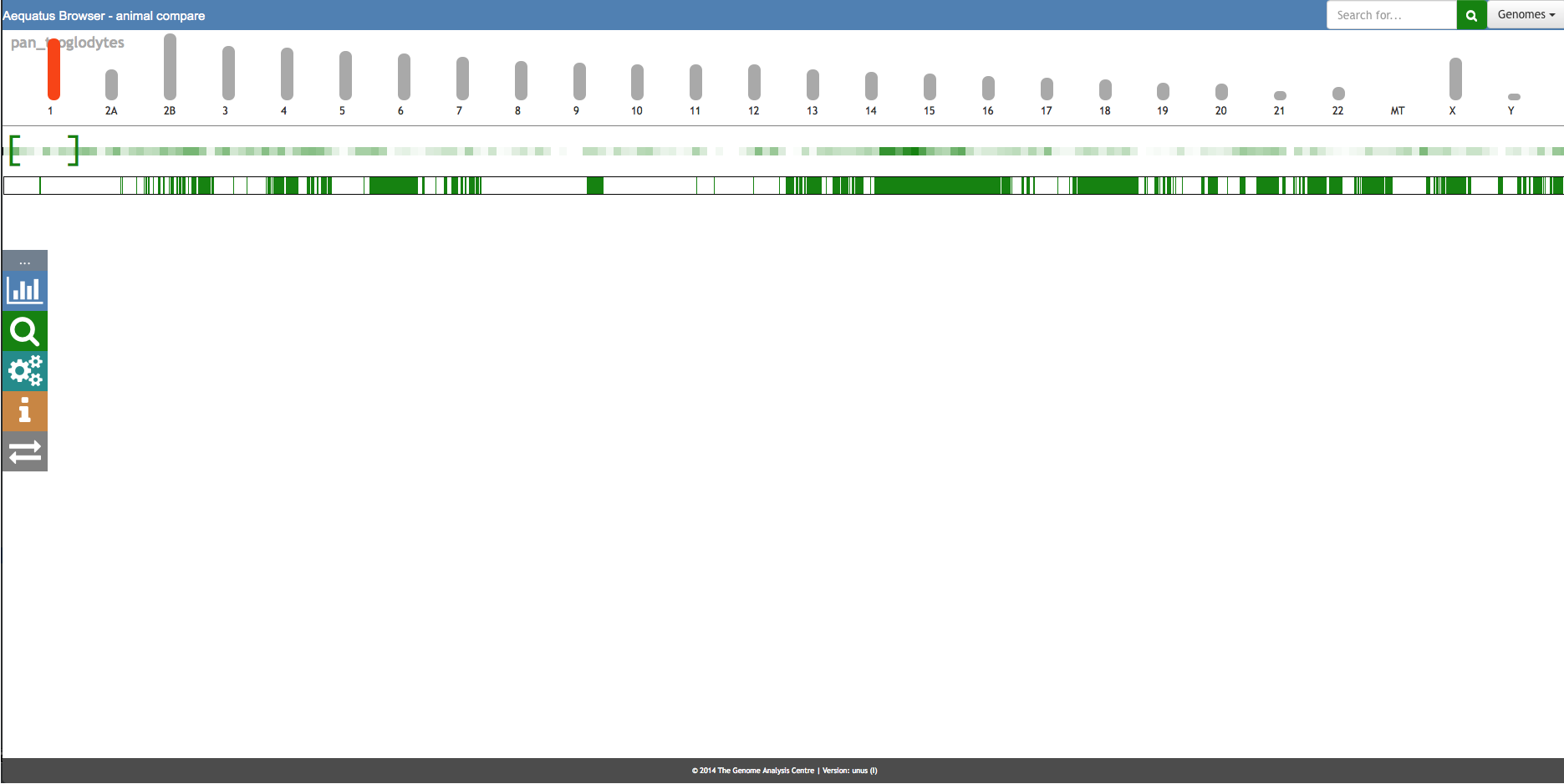
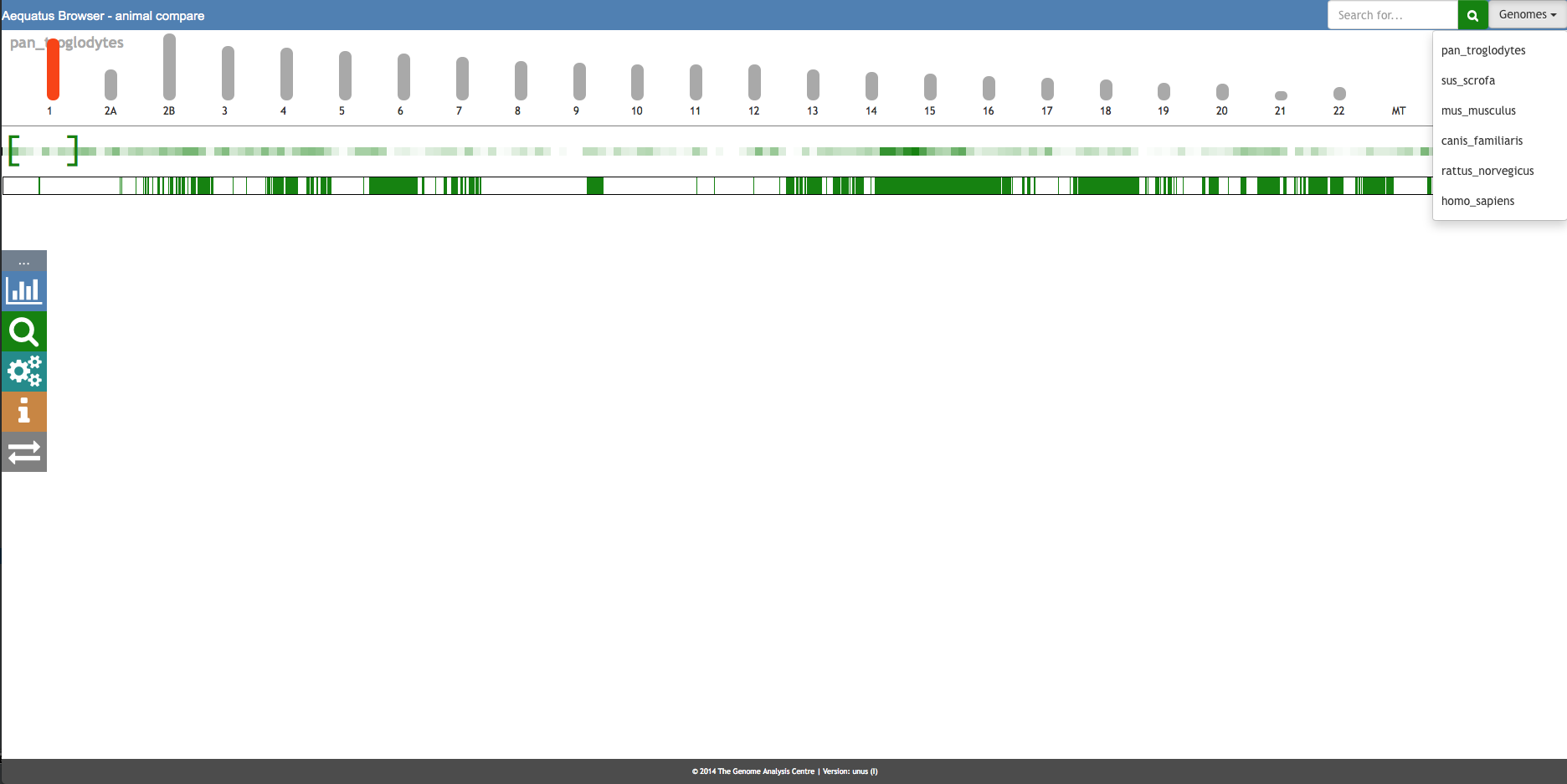
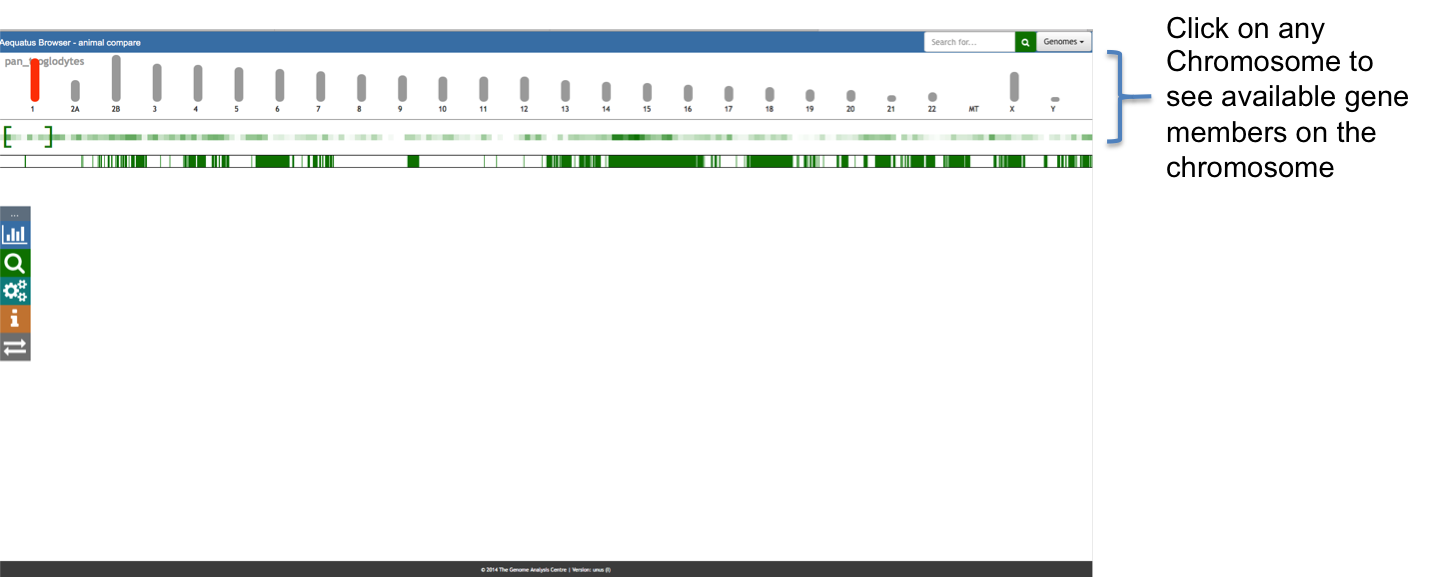
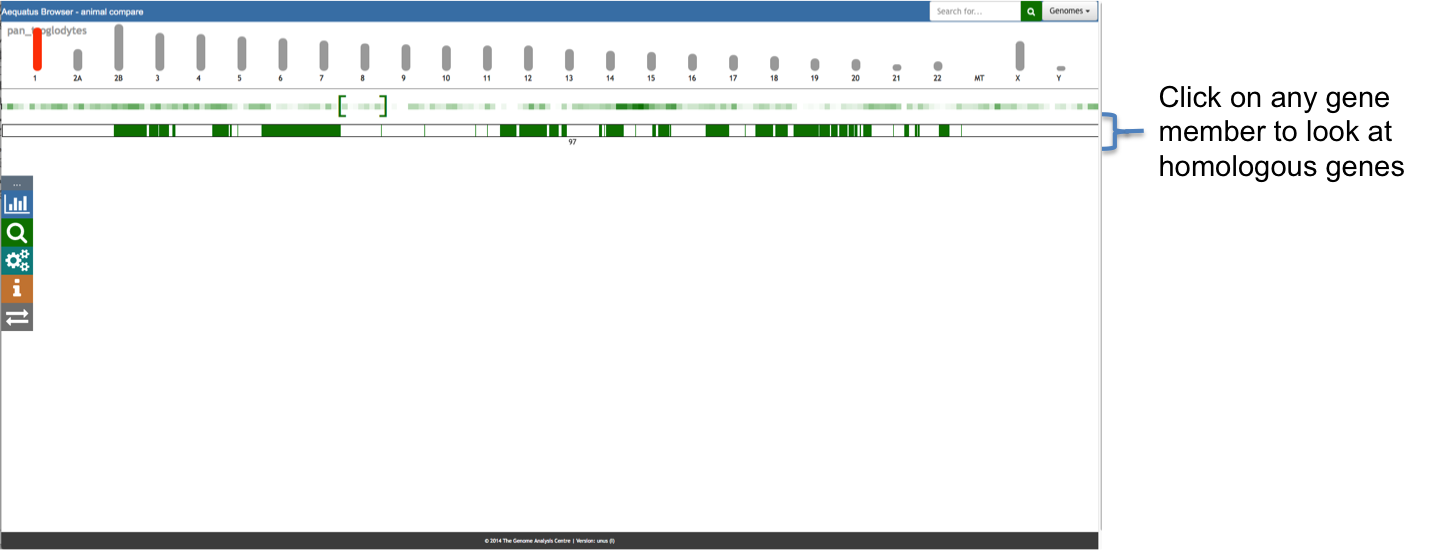
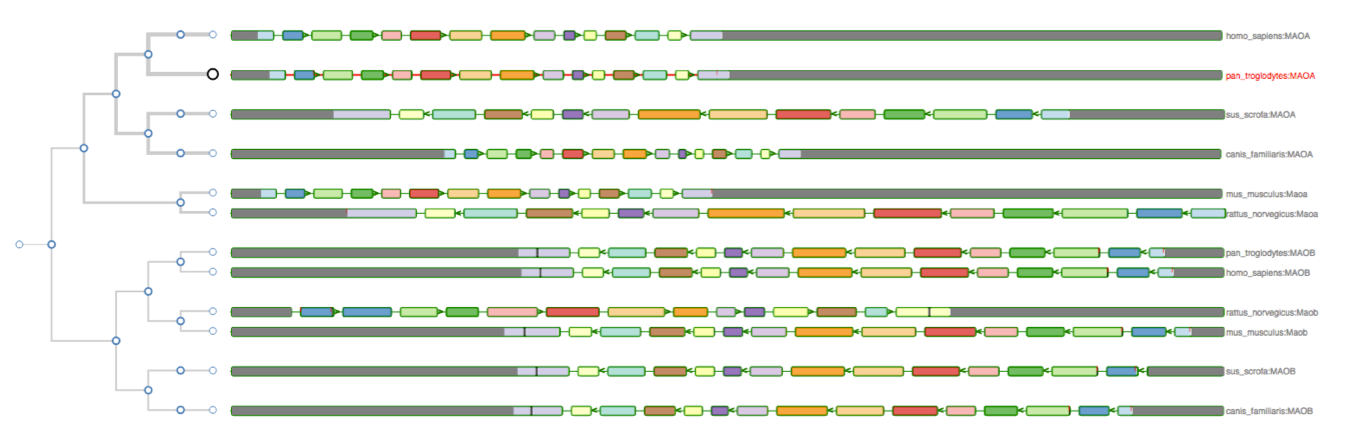
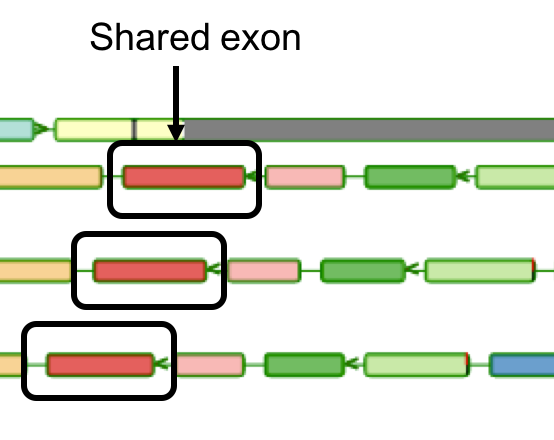
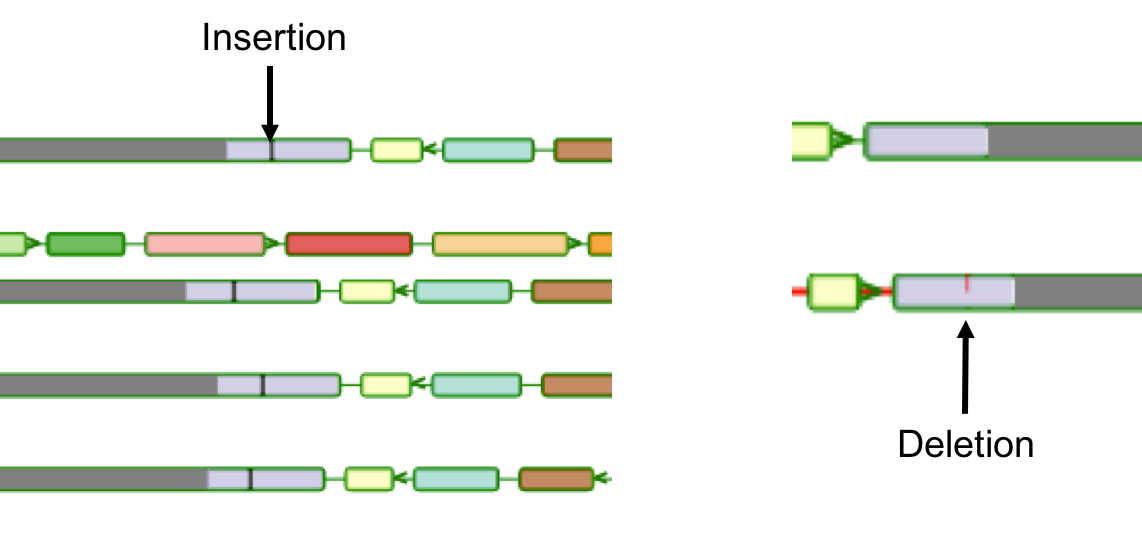
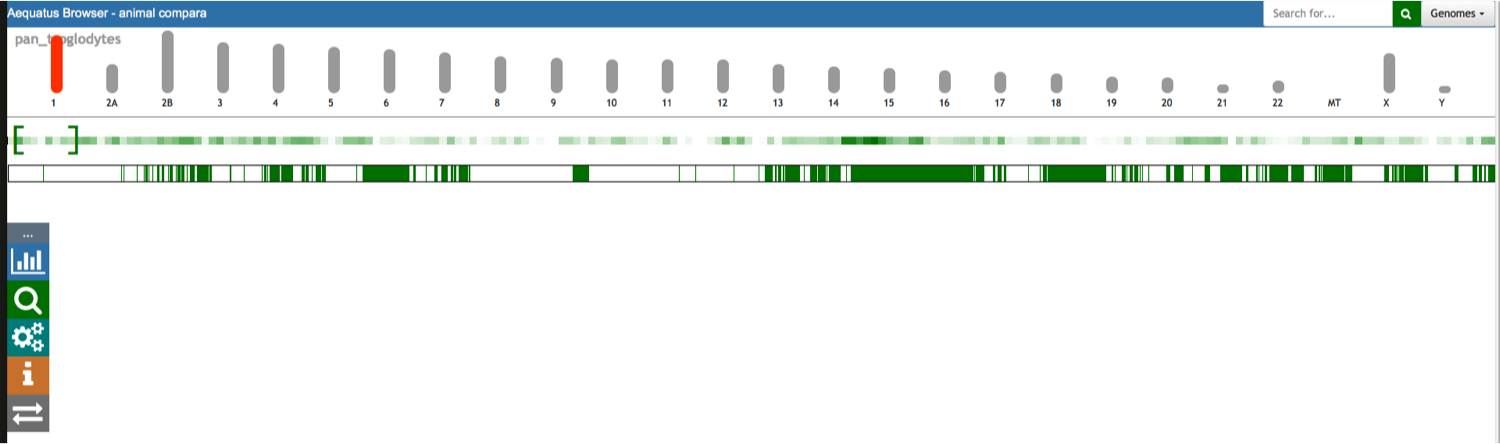
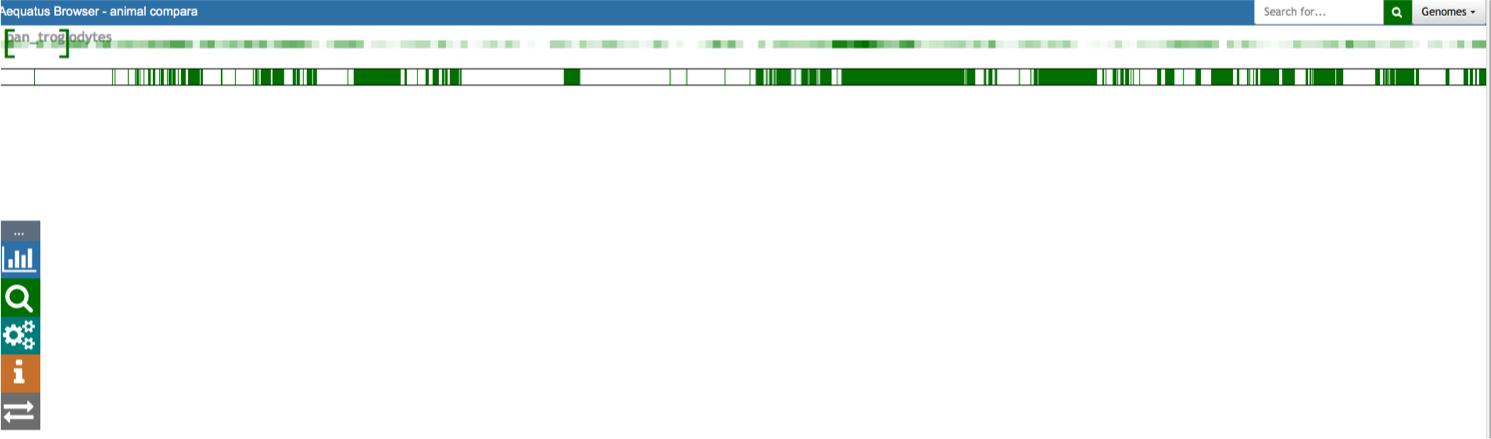
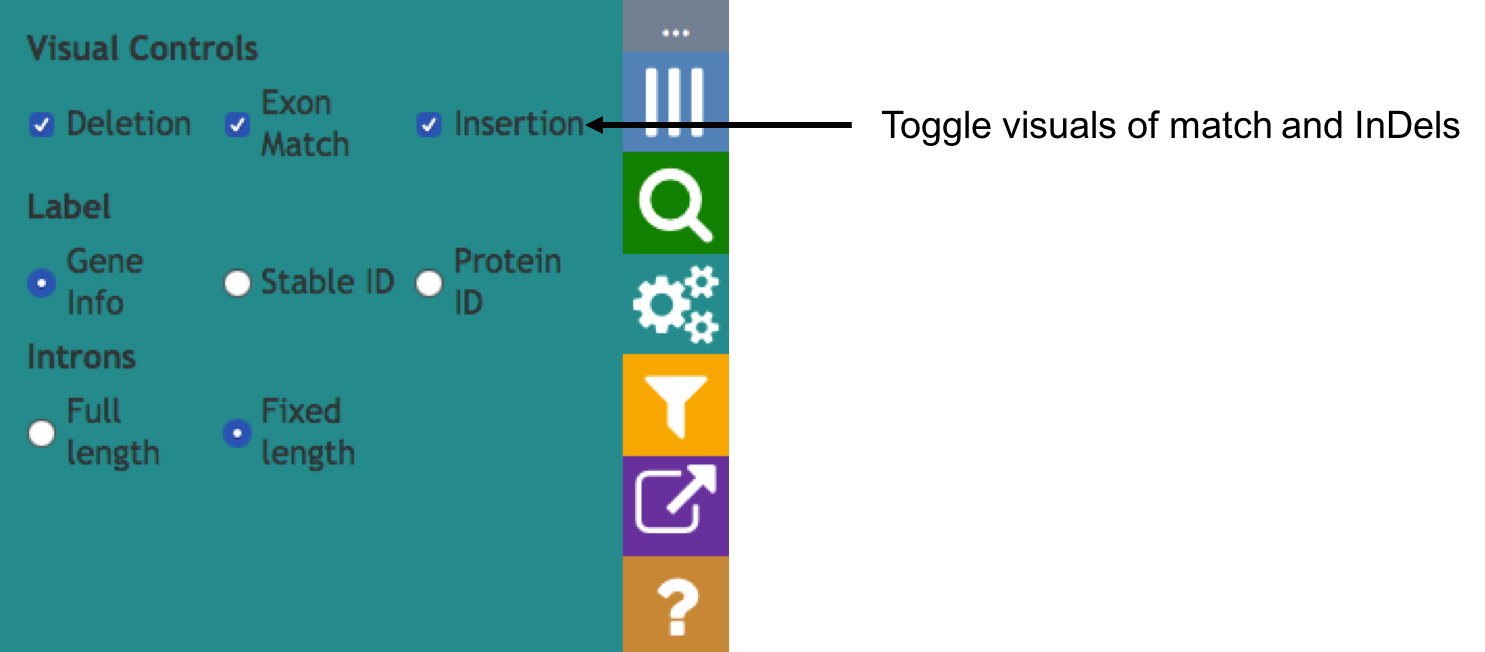
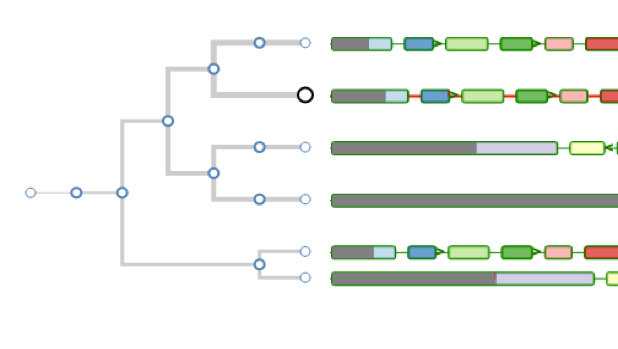
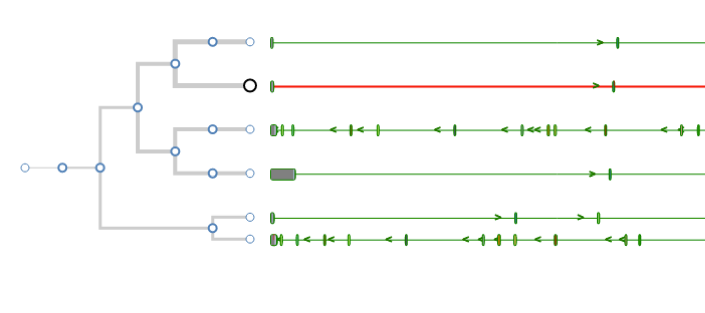
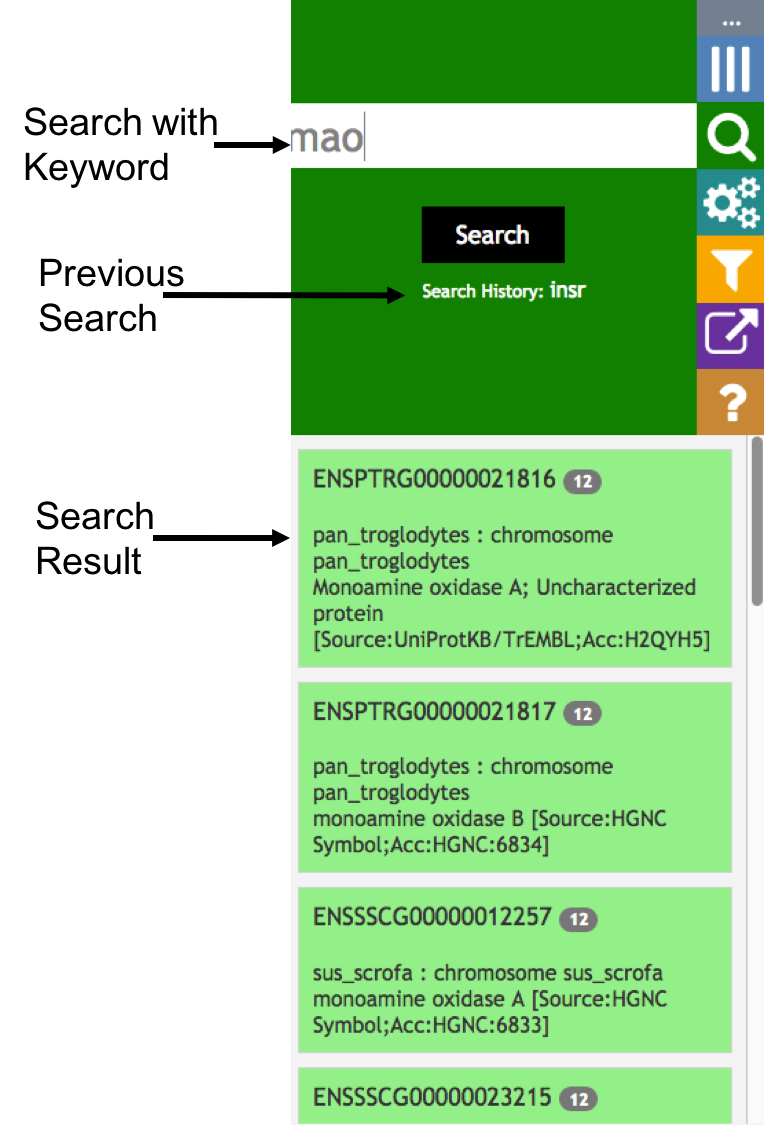
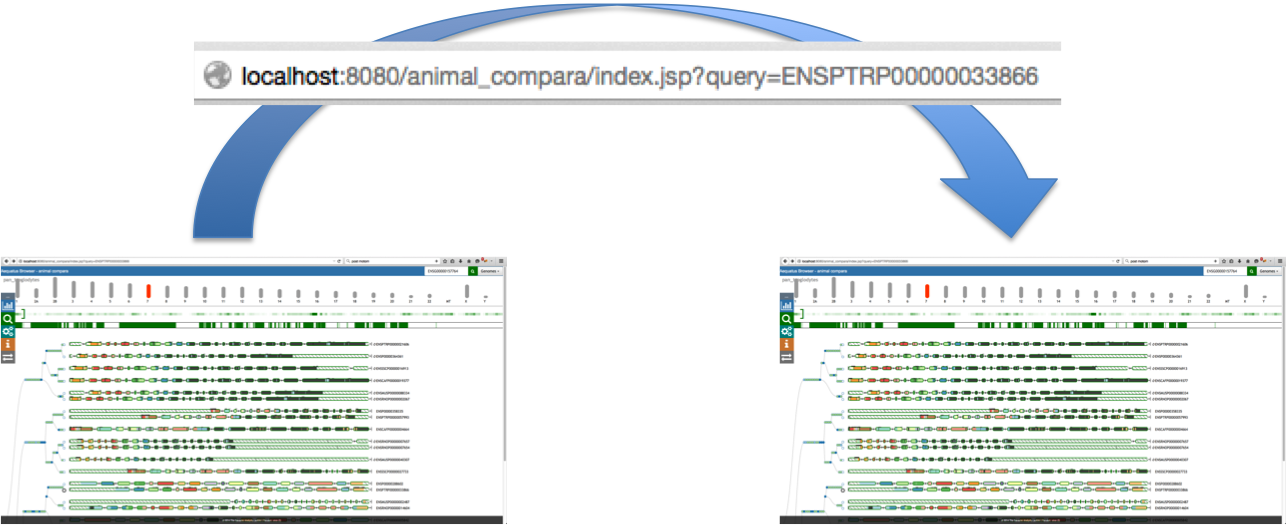
 Aequatus Browser, an open-source web-based tool developed at Earlhan Institute to visualise homologous gene structures among differing species or subtypes of a common species. Aequatus works on top of the Ensembl Compara and Core database schema. Aequatus uses precalculated gene family information and genomic alignments data in the form of CIGAR strings, from Ensembl Compara, and cross-references these sequences to Ensembl Core databases for each species to gather genomic feature information via stable_ids. Aequatus then processes the comparative and feature data to provide a visual representation of the phylogenetic and structural relationships among the set of chosen species. The ultimate goal of the Aequatus Browser is to provide a unique and informative way to render and explore complex relationships between genes from various species at a level that has so far been unrealised. Project Website: http://aequatus.earlham.ac.uk/ Source Code: https://github.com/tgac/aequatus-browser Contact: anil.thanki@earlham.ac.uk Twitter: @aequatus License: GPL V3 |